r2u: CRAN as Ubuntu Binaries
Key features
-
Full integration with
aptas every binary resolves all its dependencies: No more installations (of pre-built archives) only to discover that a shared library is missing. No more surprises. -
Full integration with
aptso that an update of a system library cannot break an R package: if a (shared) library is used by a CRAN, the package manager knows, and will not remove it. No more (R package) breakage from (system) library updates. -
Simpler and lighter than some alternatives as only run-time library packages are installed as dependencies (instead of generally heavier development packages).
-
Installations are fast, automated and reversible thanks to the package management layer.
-
Fast and well-connected mirror at r2u.stat.illinois.edu on the Internet2
-
Complete coverage with (using 24.04 for both amd64 and arm64) about 31485 CRAN packages (and 633 from BioConductor) using current versions: We support R 4.6.0, and BioConductor 3.23 (on resolute, noble and jammy; focal is in archived mode and remains at 3.21); for 24.04 (aka 'noble') and 26.04 (aka 'resolute') BioConductor is available for both amd64 and arm64.
-
Complete support for Ubuntu 20.04 ("focal"), 22.04 ("jammy"), 24.04 ("noble") and 26.04 ("resolute") on amd64, as well as 24.04 ("noble") and 26.04 ("resolute") support on arm64.
-
We update and extend (at least) the two most-current LTS releases (now 24.04 "noble" and 22.04 "jammy") daily; the previous release (20.04 "focal") has been updated up until July 2025 and remains available as is. We are currently having the 'soon-to-be released' 26.04 "resolute" in preview.
-
Fully transparent and completely open source build setup using two helper repositories r2u-builder (containing the build actions and containers) and r2u-config (with simple configuration setup).
-
Optional (but recommended) bspm use automagically connects R functions like
install.packages()toaptfor access to binaries and dependencies. -
Docker containers
rocker/r2ufrom the Rocker Project for 'focal', 'jammy', 'noble', and 'resolute'. -
GitHub Actions support to set up on Ubuntu 'latest' or via container.
Brief Demo
The gif below shows how one install.packages("tidyverse") command on an Ubuntu
20.04 system installs all packages and dependencies as binaries in 18 seconds (by passing the
R package installation to apt using bspm).
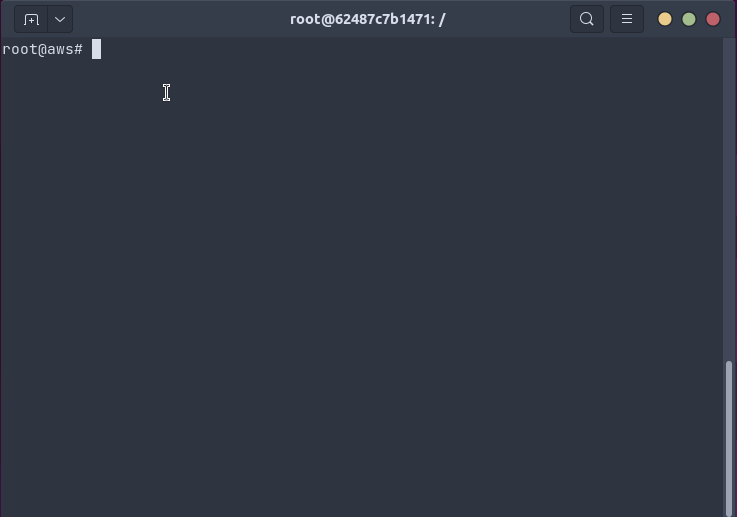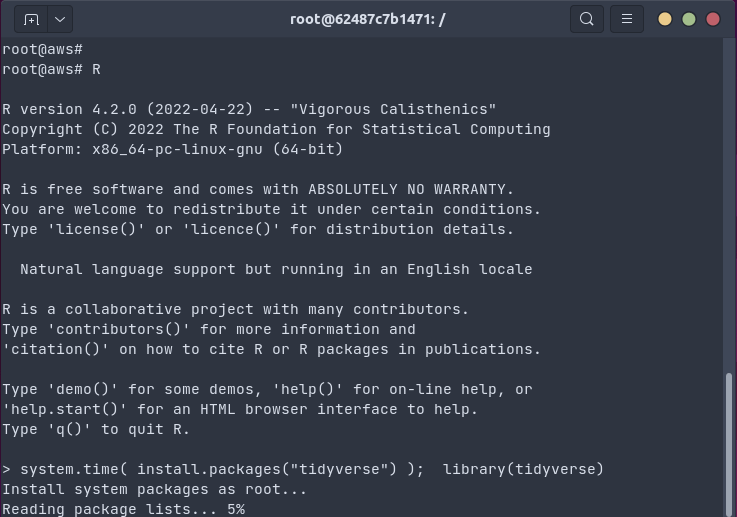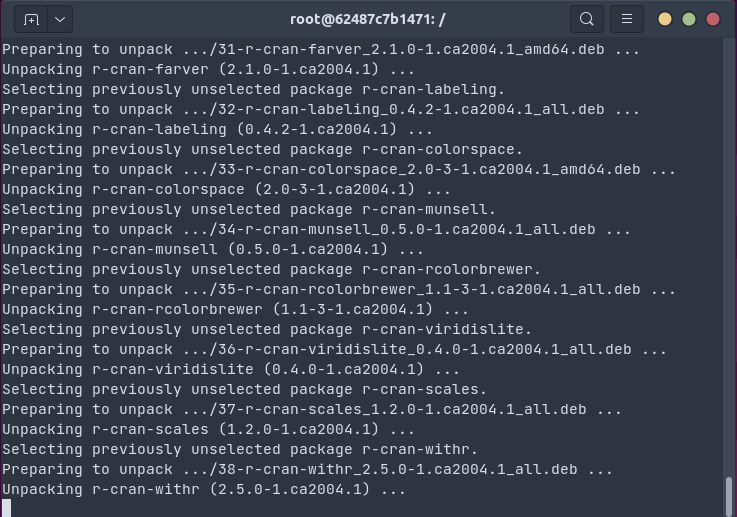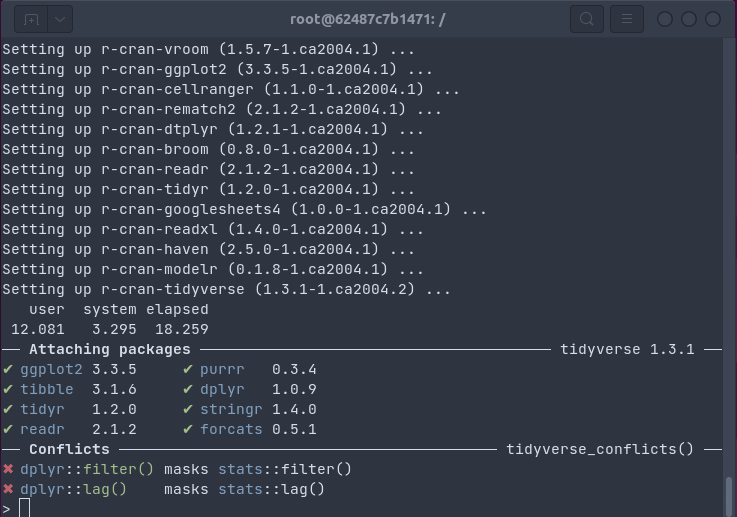
This uses the Docker container referenced below, which has been set up with the five easy setup steps detailed here.
What is Covered ?
We generally support amd64 (i.e. standard 64-bit Intel/AMD cpus, sometimes also called x86_64) for the current Ubuntu LTS release and its predecessor release, and support arm64 on the current release (more on this here). We use 'r-release' just like CRAN. So generally support (at least) the two most current LTS releases: currently 'jammy' 22.04 LTS and 'noble' 24.04 releases are fully supported with 'resolute' 26.04 ready for its upcoming release. The 'focal' 20.04 LTS release was the first one supported, and remains avaialble but (as of July 2025) no longer receives new updates.
Support for additional cpu architectures is certainly possible but somewhat unlikely due to a lack of (additional hardware) resources and time. Support for other distributions is possible but unlikely right now (due to a lack of resources and time). P3M/PPM/RSPM now appears to also support Debian which could be added at some later point.
Current versions are built under R 4.6.0 and are usable under the current R 4.6.0. BioConductor release 3.23 packages are provided when required by CRAN packages for resolute, noble and jammy; focal is at 3.21. Binaries were initially R 4.4.x based and are now built under R 4.6.0. Some older packages released when we used R 4.2.x or 4.3.x may have been built with R 4.2.x or R 4.3.x, they will still work the same with R 4.4.x as R is generally forward-compatible. There are exceptions (due to how R Core changes this) so if you notice an older package not loading please file an issue.
What is Selected ?
Everything :)
Initially, we started from cran-logs by picking the N most-downloaded packages, along with their dependencies from BioConductor. (It should be noted that for example the first 100 packages already account for approximately half the total downloads: it is a very skewed distribution.) We iterated, and fairly soon arrived of full coverage of CRAN.
So we now cover
- all CRAN packages (modulo at best a handful of blacklisted ones) including all packages needing compilation
- all BioConductor packages implied by these plus a 'healthy subset' of the highest scoring BioConductor packages (also covering e.g. all BioConductor packages in the Debian and Ubuntu distributions)
This currently results in 25706, 28188, 31485, 29106 binary packages from CRAN in "focal", "jammy", "noble", and "resolute", respectively. It also contains 431, 534, 633, and 620 BioConductor packages, respectively, from their 3.23 (resolute, noble, jammy) and 3.21 (focal) releases. (See this FAQ about why this number is higher than CRAN, and variable between releases.)
The sole exception are packages we cannot build (as we do not have the required commercial software it accessess, or do not have the required more recent toolchain component) plus a handful or so of 'odd builds' that fail and are skipped. For rust-based packages we ensure the toolchain is more recent than what the Ubuntu distribution provides so that we can build whatever is at CRAN.
What is it Based On?
Initially, for the CRAN binaries we repackaged P3M/RSPM/PPM builds (where available) or build natively. All selected BioConductor packages are built natively. For all of these, full dependency resolution and integration with the system is a key feature. For arm64 we started to build from source as we did for BioConductor. Initially we build BioConductor packages locally too; this was moved to GitHub Actions in 2025. The experience of building via GitHub Action for both arm64 and BioConductor has been generally positive so now everything is built from source at GitHub Actions in the r2u-builder repository (which used to be the r2u-bioc-builder repository).
Everything is provided as .deb binary files with proper dependency resolution by using a proper
apt repo which also has a signed Release file.
Usage and Setup
(Note that you could use one of the scripts
add_cranapt_noble.sh
(for Ubuntu 24.04), or
add_cranapt_jammy.sh
(for Ubuntu 22.04), or
add_cranapt_focal.sh
(for the older Ubuntu 20.04) to facilitate the setup. They are tested on 'empty' Ubuntu containers
of the corresponding release. However, you may prefer to execute the steps outlined here by hand.)
You can use lsb_release -cs to generate your release name: "focal", "jammy", and "noble" are
supported and you could swap "focal" or "noble" in below (or use one of the scripts).
Here, we show the setup step by step for 'jammy' aka Ubuntu 22.04 (as it is still the most-widely
used distribution per our logs, though we may update this to 24.04 soon). You should run all these
commands as root so carefully review each one. If you prefer the newer Ubuntu 24.04, please see
the
add_cranapt_noble.sh
script which also avoids the now-deprecated apt-key command).
Step 1: Update apt, install tools, fetch key
First add the repository key so that apt knows it (this is optional but recommended)
apt update -qq && apt install --yes --no-install-recommends wget \
ca-certificates gnupg
wget -q -O- https://eddelbuettel.github.io/r2u/assets/dirk_eddelbuettel_key.asc \
| tee -a /etc/apt/trusted.gpg.d/cranapt_key.asc
Step 2: Add the apt repo
Second, add the repository to the apt registry. We recommend the well-connected main mirror
provide at University of Illinois:
echo "deb [arch=amd64] https://r2u.stat.illinois.edu/ubuntu jammy main" \
> /etc/apt/sources.list.d/cranapt.list
apt update -qq
Use arch=arm64 for arm64 support (currently only available for noble).
Step 3: Ensure you have current R binaries (optional)
Third, and optionally, if you do not yet have the current R version, run these two lines (or use the standard CRAN repo setup)
wget -q -O- https://cloud.r-project.org/bin/linux/ubuntu/marutter_pubkey.asc \
| tee -a /etc/apt/trusted.gpg.d/cran_ubuntu_key.asc
echo "deb [arch=amd64] https://cloud.r-project.org/bin/linux/ubuntu jammy-cran40/" \
> /etc/apt/sources.list.d/cran_r.list
apt-key adv --keyserver keyserver.ubuntu.com --recv-keys \
67C2D66C4B1D4339 51716619E084DAB9
apt update -qq
DEBIAN_FRONTEND=noninteractive apt install --yes --no-install-recommends \
r-base-core
Use arch=arm64 for arm64 support (currently only available for noble).
Step 4: Use pinning for the r2u repo (optional)
Fourth, add repository 'pinning' as apt might get confused by some older
packages (in the Ubuntu distro) which accidentally appear with a higher
version number. See the next section for a short discussion how it ensures 'CRANapt'
sorts highest.
echo "Package: *" > /etc/apt/preferences.d/99cranapt
echo "Pin: release o=CRAN-Apt Project" >> /etc/apt/preferences.d/99cranapt
echo "Pin: release l=CRAN-Apt Packages" >> /etc/apt/preferences.d/99cranapt
echo "Pin-Priority: 700" >> /etc/apt/preferences.d/99cranapt
After that the package are known (under their r-cran-* and r-bioc-*
names). You can install them on the command-line using apt and apt-get,
via aptitude as well as other front-ends.
Step 5: Use bspm (optional)
Fifth, and also optional, install and enable the bspm
package so that the r2u (or CRANapt) as well as other R packages (available as r-*.deb binaries)
become available via install.packages() and update.packages(). Note that you may need to install
it directly from source via sudo Rscript -e 'install.packages("bspm")' to ensure it integrates
correctly with the packaging system. You should also install Python components used internally by
bspm via the sudo apt-get install
python3-{dbus,gi,apt} command.
apt install --yes --no-install-recommends python3-{dbus,gi,apt}
## Then install bspm (as root) and enable it, and enable a speed optimization
Rscript -e 'install.packages("bspm")'
RHOME=$(R RHOME)
echo "suppressMessages(bspm::enable())" >> ${RHOME}/etc/Rprofile.site
echo "options(bspm.version.check=FALSE)" >> ${RHOME}/etc/Rprofile.site
That's it! Now try it out!
About Pinning
Packages can be found in different repositories, and generally the highest available version is
the one we what---and apt picks it for us. Now, because we let apt (and related tools) pick the
packages based on versions, we may want to ensure that the CRANapt repo sorts higher than the
default repo as (older) package builds in the distribution itself may appear (to apt) to be newer
via a quirk in the sorting algorithm. A case in point was package gtable whose version in Ubuntu
was 0.3.0+dfsg-1 which accidentally sorts higher than the rebuild we made under a newer and more
consistent version number 0.3.0-1.ca2004.1.
For this issue, one possible and popular fix is to use 'apt pinning'. It can give 'higher weight' to packages from a particular repositor or tag. In the suggested example above, we give the r2u / cranapt repo a weight of 700 which is higher than the package default value of 500.
Docker
Core r2u Containers
There are also Docker containers for Ubuntu 20.04 'focal', 22.04 'jammy', 24.04 'noble', and 26.04 'resolute', respectively. Initially published as eddelbuettel/r2u, these are now also available also as rocker/r2u. They all have the features detailed above, including pinning and bspm support, already set up.
Each of the Ubuntu LTS flavors, i.e., 'focal' and 'jammy' is also available as an identical image using the release version, i.e., '20.04', '22.04', '24.04', and '26.04', respectively.
Note that with some builds of Docker (and possibly related to Ubuntu hosts) you may have to add
the --security-opt seccomp=unconfined option to your Docker invocation to take advantage of bspm
and the full system integration inside the container.
This is also documented in the FAQ.
Contributed Containers
We are now starting to see derived containers:
- BioConductor has an (alpha release) project bioc2u providing (internal ?) BioConductor builds.
- Jeffrey Girard created rstudio2u which adds RStudio to the base layer provided by r2u.
- Dirk Eddelbuettel created the r2u4ci variant for CI use (see also next section).
It is encouraging to see such specialisations based off r2u itself.
GitHub Actions
There are a number of ways to take advantage of r2u in GitHub Actions. The first, oldest, and
conceptually possibly simplest, is to switch to using the Docker container (see previous
section). This can be as easy as adding the container: statement after runs-on: in jobs:
section:
runs-on: ubuntu-latest
container:
image: rocker/r2u:latest
A complete example is provided in this action
file from R
package RcppInt64. The key advantage of this approach
is that everything for r2u is already set up (but some tools you may want CI are missing, so
please read on). This can even be used in a Matrix
setup
if one makes the container specification an argument present with, say, ubuntu-latest, but not the
others. An example is provided in this alternate actions
file which uses
the r2u4ci container variant described in the previous section. This alternate container can be
used in the preceding example, it has the advantage of bringing a handful of additional binaries
(including sudo) to the container.
An alternative approach consists of adding r2u as a step via the r2u-setup GitHub
Action:
- name: Setup r2u
uses: eddelbuettel/github-actions/r2u-setup@master
A complete example is provided in this
repo
where we use it because using the Docker container approach makes committing back via git a little
harder.
But given that r2u has also been integrated in the r-ci setup, by far the most common use likely to enable r-ci via its setup action. This is used in dozens of packages, one example is provided in this yaml file which is in fact used unchanged in many different packages.
Using r-ci via its setup action also gives the ability to
deploy the new ubuntu-slim images at GitHub which are especially useful for lighterweight task
running in under fifteen minutes (which r2u of course enables by providing binaries fast, easy and
reliably). However, initial experiments suggest that the run-time can be a little worse. This may
improve at which point this may be become more attractive. Also note that containers cannot be
launched inside ubuntu-slim so one is limited to the setup action.
Try It
Via codespaces
See the vignette Codespaces about how to launch a 'Codespace' directly in your browser, launched from the gitrepo within minutes.
This also works from your vscode installation as a remote codespace.
The vignette has more details.
Via gitpod.io
Use this link below (after possibly signing up for gitpod.io first)
and run one of the three example scripts, or just start R in the terminal window.
The gif below display running one such example to install brms from binaries in a few seconds. Using this requires only (free) GitHub and GitPod accounts.
Usage Statistics
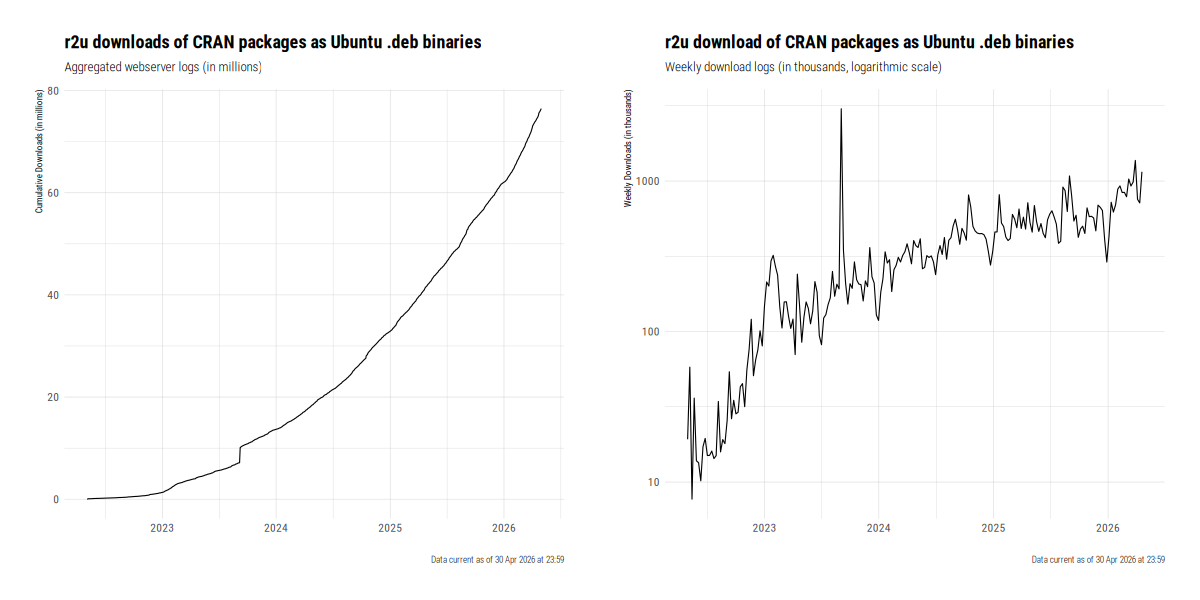
Usage is vibrant. As of spring 2026, over 4,000,000 packages are deliverd per month, with a total of now over seventy six million packages shipped. Early September 2023 also had the most recent and dramatic spike of over three million packages in two days. The following chart gives a summary of cumulative and average weekly downloads (the latter one on a log scale).
GitHub Stars
Stars are accruing at a regular pace as shown on the star history chart below.
Support
Please file issues at the GitHub issues for r2u.
Background
The (re-)recorded invited plenary talk from the 11eme Rencontres R at Mons (slides) gives some background, context and scope.
Frequently Asked Questions
Please also see the FAQ for answers to Frequently Asked Questions.
Known Issues
-
The littler package reflects build-time configuration, the RSPM/PPM binary is then expecting a different R location so it needs a binary rebuild. Added a 'force' flag, may need a list similar to the blacklist to always compiled.
-
A small number of packages do not build for lack required components; examples are ROracle and Rcplex. They, and their reverse dependencies, are blacklisted and not being built. Similarly, occassionally packages just do not build on a different cpu platform, or an older compiler.
-
r2u is an
aptrepo, which viabspmbecomes used "automagically" via standard R calls ofinstall.packages()and alike. That last part is important: package installations that do not useinstall.packages()(such asrenv,rig, ...) do not benefit frominstall.packages()callingaptfor you, and cannot take advantage of r2u viabspm. -
bspmtraces calls toinstall.packages()and maps them system-wide installation viaapt. By choice, it does not map theremove.packages()for package removal, see this issue for more discussion. Packages can be uninstalled via the system package manager using, respectively,apt,dpkgor one of graphical frontends as well as via the R functionbspm::remove_sys().
Author
Dirk Eddelbuettel
License
The repository-building code in this package is released under the GPL (>= 2).
All CRAN and BioConductor packages are released under their respective licenses.
Acknowledgment
This was made possible by the generous support of endless coffee thanks to my GitHub Sponsors.
