Codespaces
r2u for Codespaces
The .devcontainer directory provides a small self-containted file devcontainer.json to
launch an executable environment R using r2u. It is based on the example in
Grant McDermott's
codespaces-r2u repo and
reuses its documentation. It is driven by the Rocker Project's Devcontainer
Features repo
creating a fully functioning R environment for cloud use in a few minutes.
And thanks to r2u you can add easily to this environment by installing
new R packages in a fast and failsafe way.
Try it out
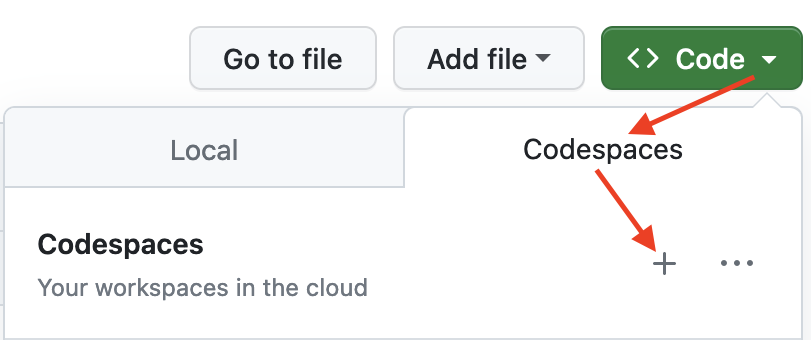
To get started, simply click on the green "Code" button at the top right. Then select the "Codespaces" tab and click the "+" symbol to start a new Codespace.
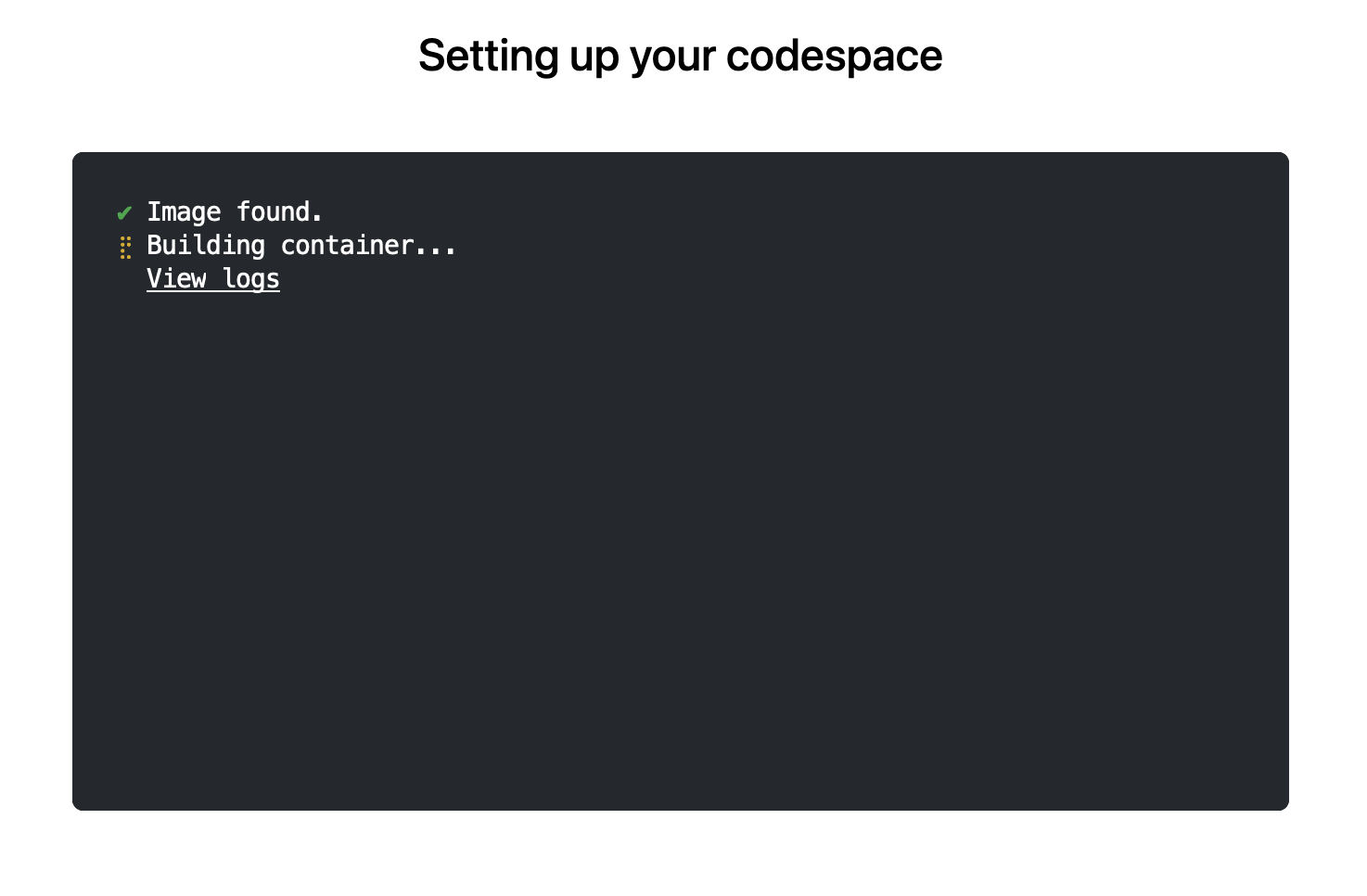
The first time you do this, it will open up a new browser tab where your Codespace is being instantiated. This first-time instantiation will take a few minutes (feel free to click "View logs" to see how things are progressing) so please be patient. Once built, your Codespace will deploy almost immediately when you use it again in the future.
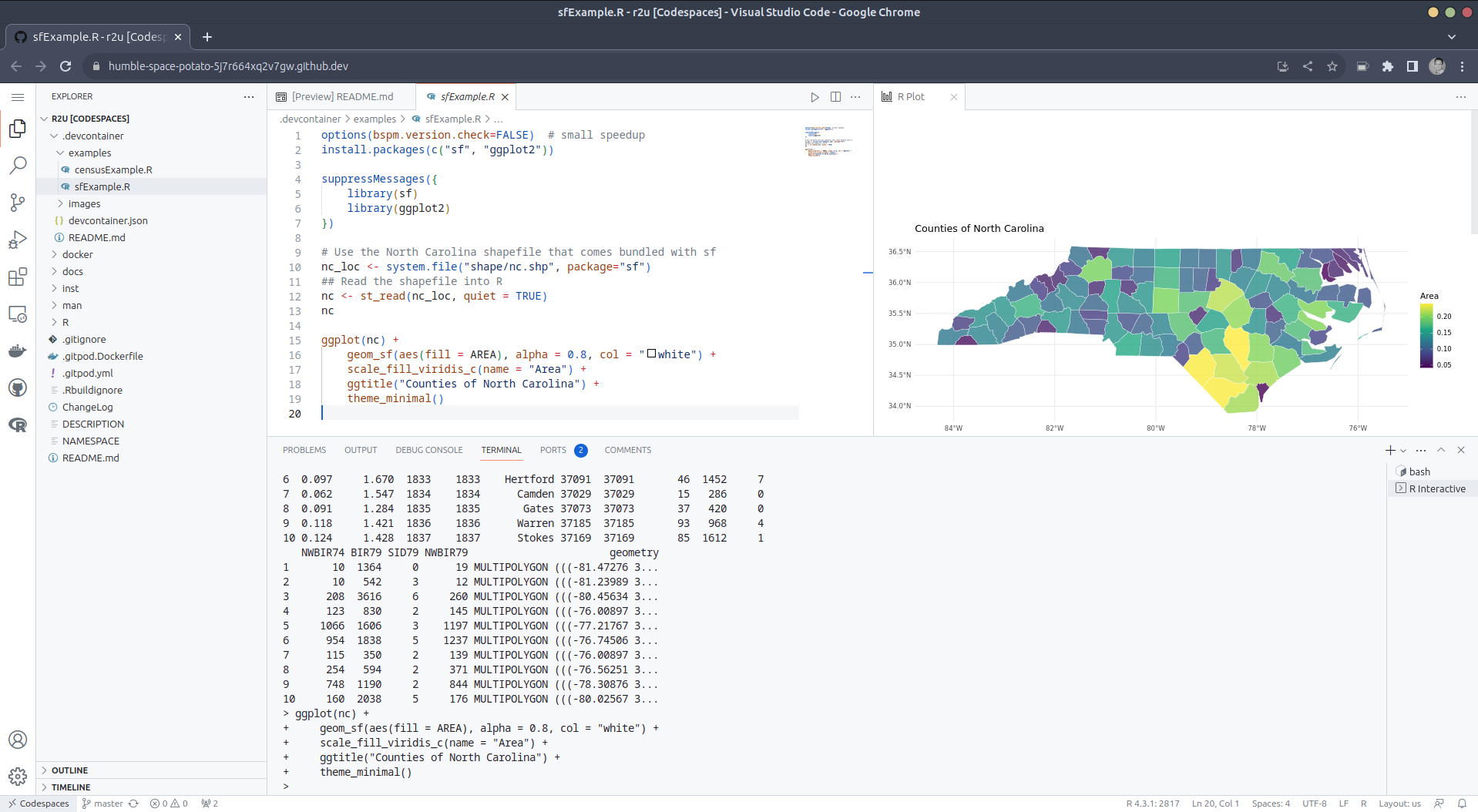
After the VS Code editor opens up in your browser, feel free to open up the
examples/sfExample.R file. It demonstrates how
r2u enables us install packages and their system-dependencies with
ease, here installing packages sf (including all its geospatial
dependencies) and ggplot2 (including all its dependencies). You
can run the code easily in the browser environment: Highlight or hover over
line(s) and execute them by hitting Cmd+Return (Mac) / Ctrl+Return
(Linux / Windows).
Do not forget to close your Codespace once you have finished using it. Click the "Codespaces" tab at the very bottom left of your code editor / browser and select "Close Current Codespace" in the resulting pop-up box. You can restart it at any time, for example by going to https://github.com/codespaces and clicking on your instance.
Extend r2u with r-universe
r2u offers "fast, easy, reliable" access to all of CRAN via binaries
for Ubuntu focal and jammy. When using the latter (as is the default), it
can be combined with r-universe and its Ubuntu jammy binaries.
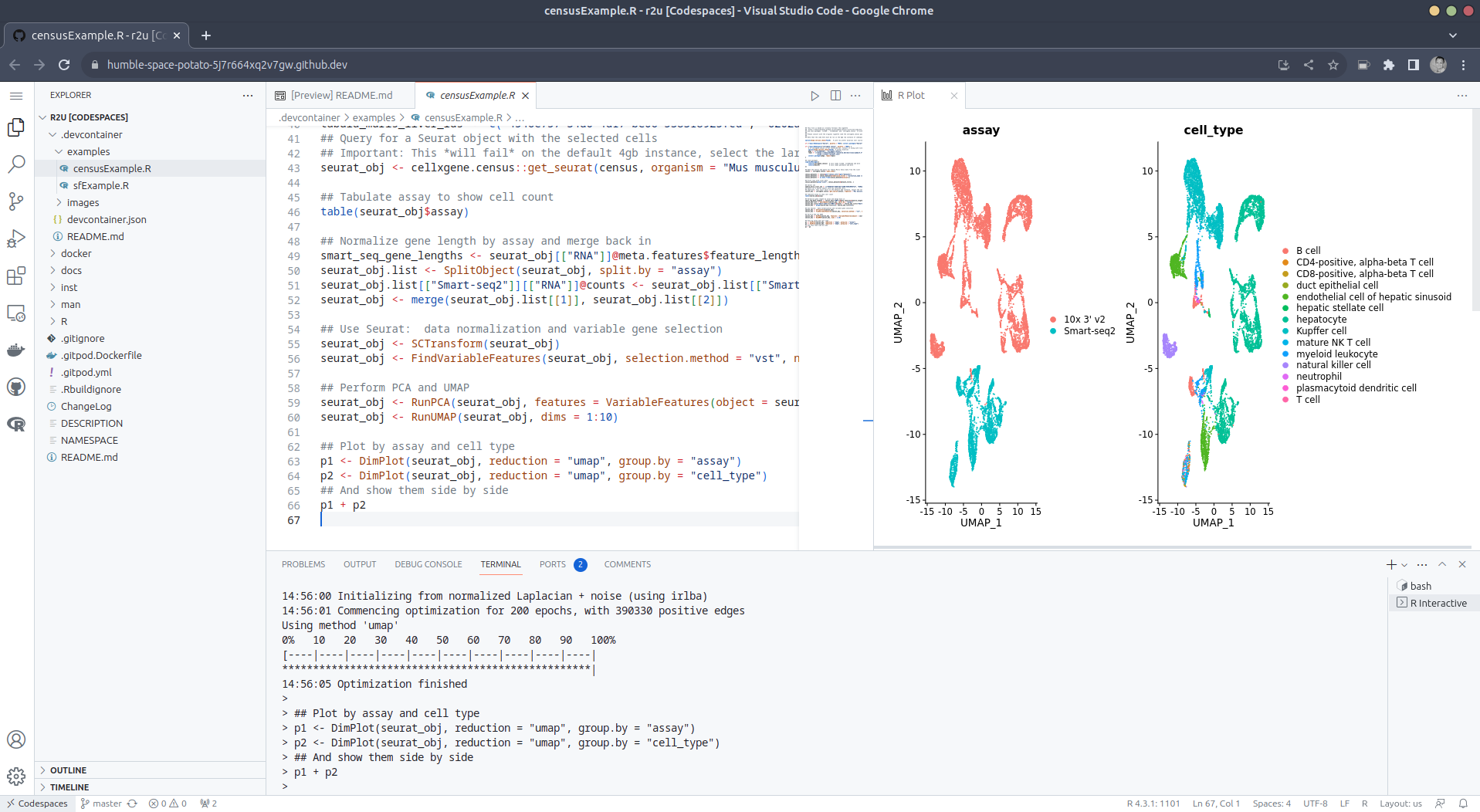
We demontrates this in a second example file
examples/censusExample.R
which install both the
cellxgene-census and
tiledbsoma R packages as
binaries from r-universe (along with about 100 dependencies),
downloads single-cell data from Census and uses
Seurat to create PCA and UMAP
decomposition plots. Note that in order run this you have to change the
Codespaces default instance from 'small' (4gb ram) to 'large' (16gb ram).
Local DevContainer build
Codespaces are DevContainers running in the cloud (where DevContainers are
themselves just Docker images running with some VS Code sugar on top). This
gives you the very powerful ability to 'edit locally' but 'run remotely' in
the hosted codespace. To test this setup locally, simply clone the repo and
open it up in VS Code. You will need to have Docker installed and running on
your system (see here). You will
also need the Remote Development extension
(you will probably be prompted to install it automatically if you do not have
it yet). Select "Reopen in Container" when prompted. Otherwise, click the
>< tab at the very bottom left of your VS Code editor and select this
option. To shut down the container, simply click the same button and choose
"Reopen Folder Locally". You can always search for these commands via the
command palette too (Cmd+Shift+p / Ctrl+Shift+p).
Use in Your Repo
To add this ability of launching Codespaces in the browser (or editor) to a repo of yours, create a
directory .devcontainers in your selected repo, and add the file
.devcontainers/devcontainer.json. You can customize it by
enabling other feature, or use the postCreateCommand field to install packages (while taking full
advantage of r2u).
Acknowledgments
There are a few key "plumbing" pieces that make everything work here. Thanks to:
- My Rocker Project colleague @eitsupi for maintaining the R DevContainer Features.
- @renkun-ken and the rest of the VS Code R extension team.
- @Enchufa2 for
bspmmaking package installation to the sysstem so seamless. - @grantmcdermott for the initial codespaces-r2u setup from which we derived this.
- Last but not least everybody who helped me make r2u possible, tested it, or sent hints for improvement.